Spaces:
Running
on
Zero
Running
on
Zero
Upload 7 files
Browse files- Model_Class.py +109 -0
- Model_Seg.py +99 -0
- __init__.py +0 -0
- anatomy_aware_pipeline.png +0 -0
- app.py +133 -0
- requirements_small.txt +12 -0
- utils.py +353 -0
Model_Class.py
ADDED
|
@@ -0,0 +1,109 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
import pytorch_lightning as pl
|
| 2 |
+
import torch
|
| 3 |
+
import torch.nn as nn
|
| 4 |
+
import utils
|
| 5 |
+
from torchvision.models import resnet50
|
| 6 |
+
import torch
|
| 7 |
+
from monai.transforms import (
|
| 8 |
+
Compose, Resize, ResizeWithPadOrCrop,
|
| 9 |
+
)
|
| 10 |
+
from pytorch_grad_cam import GradCAM
|
| 11 |
+
import matplotlib.colors as mcolors
|
| 12 |
+
import matplotlib.pyplot as plt
|
| 13 |
+
import numpy as np
|
| 14 |
+
from PIL import Image
|
| 15 |
+
from io import BytesIO
|
| 16 |
+
|
| 17 |
+
class ResNet(pl.LightningModule):
|
| 18 |
+
def __init__(self):
|
| 19 |
+
super().__init__()
|
| 20 |
+
self.save_hyperparameters()
|
| 21 |
+
|
| 22 |
+
|
| 23 |
+
backbone = resnet50()
|
| 24 |
+
num_input_channel = 1
|
| 25 |
+
layer = backbone.conv1
|
| 26 |
+
new_layer = nn.Conv2d(
|
| 27 |
+
in_channels=num_input_channel,
|
| 28 |
+
out_channels=layer.out_channels,
|
| 29 |
+
kernel_size=layer.kernel_size,
|
| 30 |
+
stride=layer.stride,
|
| 31 |
+
padding=layer.padding,
|
| 32 |
+
bias=layer.bias,
|
| 33 |
+
)
|
| 34 |
+
new_layer.weight = nn.Parameter(layer.weight.sum(dim=1, keepdim=True))
|
| 35 |
+
|
| 36 |
+
backbone.conv1 = new_layer
|
| 37 |
+
|
| 38 |
+
|
| 39 |
+
backbone.fc = nn.Sequential(
|
| 40 |
+
nn.Linear(2048, 1024),
|
| 41 |
+
nn.ReLU(),
|
| 42 |
+
nn.BatchNorm1d(1024),
|
| 43 |
+
nn.Dropout(0),
|
| 44 |
+
nn.Linear(1024, 2),
|
| 45 |
+
)
|
| 46 |
+
|
| 47 |
+
self.model = backbone
|
| 48 |
+
|
| 49 |
+
def forward(self, x):
|
| 50 |
+
out = self.model(x)
|
| 51 |
+
return out
|
| 52 |
+
|
| 53 |
+
|
| 54 |
+
val_transforms_416x628 = Compose(
|
| 55 |
+
[
|
| 56 |
+
utils.CustomCLAHE(),
|
| 57 |
+
Resize(spatial_size=628, mode="bilinear", align_corners=True, size_mode="longest"),
|
| 58 |
+
ResizeWithPadOrCrop(spatial_size=(416, 628)),
|
| 59 |
+
]
|
| 60 |
+
)
|
| 61 |
+
|
| 62 |
+
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
|
| 63 |
+
checkpoint = torch.load("classification_model.ckpt", map_location=torch.device('cpu'))
|
| 64 |
+
model = ResNet().to(device)
|
| 65 |
+
model.load_state_dict(checkpoint["state_dict"])
|
| 66 |
+
model.eval()
|
| 67 |
+
|
| 68 |
+
|
| 69 |
+
def load_and_classify_image(image_path):
|
| 70 |
+
image = val_transforms_416x628(image_path)
|
| 71 |
+
image = image.unsqueeze(0).to(device)
|
| 72 |
+
|
| 73 |
+
with torch.no_grad():
|
| 74 |
+
prediction = model(image)
|
| 75 |
+
prediction = torch.nn.functional.softmax(prediction, dim=1).squeeze(0)
|
| 76 |
+
return prediction.to('cpu'), image.to('cpu')
|
| 77 |
+
|
| 78 |
+
|
| 79 |
+
def make_GradCAM(image):
|
| 80 |
+
|
| 81 |
+
model.eval()
|
| 82 |
+
target_layers = [model.model.layer4[-1]]
|
| 83 |
+
|
| 84 |
+
arr = image.numpy().squeeze()
|
| 85 |
+
cam = GradCAM(model=model, target_layers=target_layers)
|
| 86 |
+
targets = None
|
| 87 |
+
grayscale_cam = cam(
|
| 88 |
+
input_tensor=image,
|
| 89 |
+
targets=targets,
|
| 90 |
+
aug_smooth=False,
|
| 91 |
+
eigen_smooth=True,
|
| 92 |
+
)
|
| 93 |
+
grayscale_cam = grayscale_cam.squeeze()
|
| 94 |
+
|
| 95 |
+
jet = plt.colormaps.get_cmap("inferno")
|
| 96 |
+
newcolors = jet(np.linspace(0, 1, 256))
|
| 97 |
+
newcolors[0, :3] = 0
|
| 98 |
+
new_jet = mcolors.ListedColormap(newcolors)
|
| 99 |
+
|
| 100 |
+
plt.figure(figsize=(10, 10))
|
| 101 |
+
plt.imshow(arr, cmap='gray')
|
| 102 |
+
plt.imshow(grayscale_cam, cmap=new_jet, alpha=0.5)
|
| 103 |
+
plt.axis('off')
|
| 104 |
+
buffer2 = BytesIO()
|
| 105 |
+
plt.savefig(buffer2, format='png', bbox_inches='tight', pad_inches=0)
|
| 106 |
+
buffer2.seek(0)
|
| 107 |
+
gradcam_image = np.array(Image.open(buffer2)).squeeze()
|
| 108 |
+
|
| 109 |
+
return gradcam_image
|
Model_Seg.py
ADDED
|
@@ -0,0 +1,99 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
import torch
|
| 2 |
+
import numpy as np
|
| 3 |
+
from monai.transforms import Compose, LoadImage, EnsureChannelFirst, Lambda, Resize, NormalizeIntensity, GaussianSmooth, ScaleIntensity, AsDiscrete, KeepLargestConnectedComponent, Invert, Rotate90, SaveImage, Transform
|
| 4 |
+
from monai.inferers import SlidingWindowInferer
|
| 5 |
+
from monai.networks.nets import UNet
|
| 6 |
+
|
| 7 |
+
class RgbaToGrayscale(Transform):
|
| 8 |
+
def __call__(self, x):
|
| 9 |
+
# squeeze last dimension, to ensure C, H, W format
|
| 10 |
+
x = x.squeeze(-1)
|
| 11 |
+
# Ensure the tensor is 3D (channels, height, width)
|
| 12 |
+
if x.ndim != 3:
|
| 13 |
+
raise ValueError(f"Input tensor must be 3D. Shape: {x.shape}")
|
| 14 |
+
|
| 15 |
+
# Check the number of channels
|
| 16 |
+
if x.shape[0] == 4: # Assuming RGBA
|
| 17 |
+
rgb_weights = torch.tensor([0.2989, 0.5870, 0.1140], device=x.device)
|
| 18 |
+
# Apply weights to RGB channels, output should retain one channel dimension
|
| 19 |
+
grayscale = torch.einsum('cwh,c->wh', x[:3, :, :], rgb_weights).unsqueeze(0)
|
| 20 |
+
elif x.shape[0] == 3: # Assuming RGB
|
| 21 |
+
rgb_weights = torch.tensor([0.2989, 0.5870, 0.1140], device=x.device)
|
| 22 |
+
grayscale = torch.einsum('cwh,c->wh', x, rgb_weights).unsqueeze(0)
|
| 23 |
+
elif x.shape[0] == 1: # Already grayscale
|
| 24 |
+
grayscale = x
|
| 25 |
+
else:
|
| 26 |
+
raise ValueError(f"Unsupported channel number: {x.shape[0]}")
|
| 27 |
+
return grayscale
|
| 28 |
+
|
| 29 |
+
def inverse(self, x):
|
| 30 |
+
# Simply return the input as the output
|
| 31 |
+
return x
|
| 32 |
+
|
| 33 |
+
model = UNet(
|
| 34 |
+
spatial_dims=2,
|
| 35 |
+
in_channels=1,
|
| 36 |
+
out_channels=4,
|
| 37 |
+
channels=[64, 128, 256, 512],
|
| 38 |
+
strides=[2, 2, 2],
|
| 39 |
+
num_res_units=3
|
| 40 |
+
)
|
| 41 |
+
|
| 42 |
+
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
|
| 43 |
+
|
| 44 |
+
checkpoint_path = 'segmentation_model.pt'
|
| 45 |
+
checkpoint = torch.load(checkpoint_path, map_location=device)
|
| 46 |
+
assert model.state_dict().keys() == checkpoint['network'].keys(), "Model and checkpoint keys do not match"
|
| 47 |
+
|
| 48 |
+
model.load_state_dict(checkpoint['network'])
|
| 49 |
+
model.eval()
|
| 50 |
+
|
| 51 |
+
# Define transforms for preprocessing
|
| 52 |
+
pre_transforms = Compose([
|
| 53 |
+
LoadImage(image_only=True),
|
| 54 |
+
EnsureChannelFirst(),
|
| 55 |
+
RgbaToGrayscale(), # Convert RGBA to grayscale
|
| 56 |
+
Resize(spatial_size=(768, 768)),
|
| 57 |
+
Lambda(func=lambda x: x.squeeze(-1)), # Adjust if the input image has an extra unwanted dimension
|
| 58 |
+
NormalizeIntensity(),
|
| 59 |
+
GaussianSmooth(sigma=0.1),
|
| 60 |
+
ScaleIntensity(minv=-1, maxv=1)
|
| 61 |
+
])
|
| 62 |
+
|
| 63 |
+
|
| 64 |
+
|
| 65 |
+
# Define transforms for postprocessing
|
| 66 |
+
post_transforms = Compose([
|
| 67 |
+
AsDiscrete(argmax=True, to_onehot=4),
|
| 68 |
+
KeepLargestConnectedComponent(),
|
| 69 |
+
AsDiscrete(argmax=True),
|
| 70 |
+
Invert(pre_transforms),
|
| 71 |
+
#SaveImage(output_dir='./', output_postfix='seg', output_ext='.nii', resample=False)
|
| 72 |
+
])
|
| 73 |
+
|
| 74 |
+
|
| 75 |
+
|
| 76 |
+
def load_and_segment_image(input_image_path):
|
| 77 |
+
|
| 78 |
+
|
| 79 |
+
image_tensor = pre_transforms(input_image_path)
|
| 80 |
+
image_tensor = image_tensor.unsqueeze(0).to(device)
|
| 81 |
+
|
| 82 |
+
# Inference using SlidingWindowInferer
|
| 83 |
+
inferer = SlidingWindowInferer(roi_size=(512, 512), sw_batch_size=16, overlap=0.75)
|
| 84 |
+
with torch.no_grad():
|
| 85 |
+
outputs = inferer(image_tensor, model)
|
| 86 |
+
|
| 87 |
+
|
| 88 |
+
outputs = outputs.squeeze(0)
|
| 89 |
+
|
| 90 |
+
processed_outputs = post_transforms(outputs)
|
| 91 |
+
|
| 92 |
+
# rotate
|
| 93 |
+
rotate = Rotate90(spatial_axes=(0, 1), k=3)
|
| 94 |
+
processed_outputs = rotate(processed_outputs).to('cpu')
|
| 95 |
+
|
| 96 |
+
output_array = processed_outputs.squeeze().detach().numpy().astype(np.uint8)
|
| 97 |
+
|
| 98 |
+
|
| 99 |
+
return output_array
|
__init__.py
ADDED
|
File without changes
|
anatomy_aware_pipeline.png
ADDED
|
app.py
ADDED
|
@@ -0,0 +1,133 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
import gradio as gr
|
| 2 |
+
import utils
|
| 3 |
+
import Model_Class
|
| 4 |
+
import Model_Seg
|
| 5 |
+
|
| 6 |
+
import SimpleITK as sitk
|
| 7 |
+
import torch
|
| 8 |
+
from numpy import uint8
|
| 9 |
+
import spaces
|
| 10 |
+
image_base64 = utils.image_to_base64("anatomy_aware_pipeline.png")
|
| 11 |
+
article_html = f"<img src='data:image/png;base64,{image_base64}' alt='Anatomical pipeline illustration' style='width:100%;'>"
|
| 12 |
+
|
| 13 |
+
description_markdown = """
|
| 14 |
+
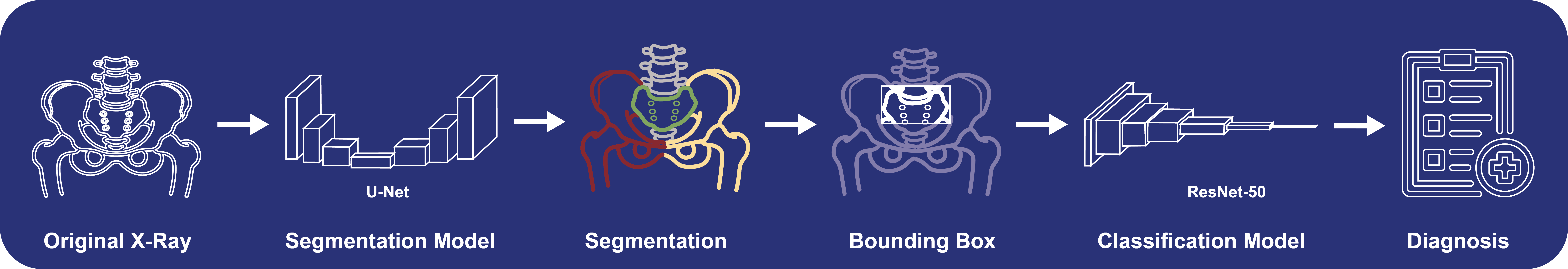
- This tool combines a U-Net Segmentation Model with a ResNet-50 for Classification.
|
| 15 |
+
- **Usage:** Just drag a pelvic x-ray into the box and hit run.
|
| 16 |
+
- **Process:** The input image will be segmented and cropped to the SIJ before classification.
|
| 17 |
+
- **Please Note:** This tool is intended for research purposes only.
|
| 18 |
+
- **Privacy:** This tool runs completely locally, ensuring data privacy.
|
| 19 |
+
"""
|
| 20 |
+
|
| 21 |
+
css = """
|
| 22 |
+
|
| 23 |
+
h1 {
|
| 24 |
+
text-align: center;
|
| 25 |
+
display:block;
|
| 26 |
+
}
|
| 27 |
+
.markdown-block {
|
| 28 |
+
background-color: #0b0f1a; /* Light gray background */
|
| 29 |
+
color: black; /* Black text */
|
| 30 |
+
padding: 10px; /* Padding around the text */
|
| 31 |
+
border-radius: 5px; /* Rounded corners */
|
| 32 |
+
box-shadow: 0 0 10px rgba(11,15,26,1);
|
| 33 |
+
display: inline-flex; /* Use inline-flex to shrink to content size */
|
| 34 |
+
flex-direction: column;
|
| 35 |
+
justify-content: center; /* Vertically center content */
|
| 36 |
+
align-items: center; /* Horizontally center items within */
|
| 37 |
+
margin: auto; /* Center the block */
|
| 38 |
+
}
|
| 39 |
+
|
| 40 |
+
.markdown-block ul, .markdown-block ol {
|
| 41 |
+
background-color: #1e2936;
|
| 42 |
+
border-radius: 5px;
|
| 43 |
+
padding: 10px;
|
| 44 |
+
box-shadow: 0 0 10px rgba(0,0,0,0.3);
|
| 45 |
+
padding-left: 20px; /* Adjust padding for bullet alignment */
|
| 46 |
+
text-align: left; /* Ensure text within list is left-aligned */
|
| 47 |
+
list-style-position: inside;/* Ensures bullets/numbers are inside the content flow */
|
| 48 |
+
}
|
| 49 |
+
|
| 50 |
+
footer {
|
| 51 |
+
display:none !important
|
| 52 |
+
}
|
| 53 |
+
"""
|
| 54 |
+
|
| 55 |
+
@spaces.GPU
|
| 56 |
+
def predict_image(input_image, input_file):
|
| 57 |
+
|
| 58 |
+
if input_image is not None:
|
| 59 |
+
image_path = input_image
|
| 60 |
+
|
| 61 |
+
elif input_file is not None:
|
| 62 |
+
image_path = input_file
|
| 63 |
+
|
| 64 |
+
else:
|
| 65 |
+
return None , None , "Please input an image before pressing run" , None , None
|
| 66 |
+
|
| 67 |
+
image_mask = Model_Seg.load_and_segment_image(image_path)
|
| 68 |
+
|
| 69 |
+
overlay_image_np, original_image_np = utils.overlay_mask(image_path, image_mask)
|
| 70 |
+
|
| 71 |
+
image_mask_im = sitk.GetImageFromArray(image_mask[None, :, :].astype(uint8))
|
| 72 |
+
image_im = sitk.GetImageFromArray(original_image_np[None, :, :].astype(uint8))
|
| 73 |
+
cropped_boxed_im, _ = utils.mask_and_crop(image_im, image_mask_im)
|
| 74 |
+
|
| 75 |
+
cropped_boxed_array = sitk.GetArrayFromImage(cropped_boxed_im)
|
| 76 |
+
cropped_boxed_array_disp = cropped_boxed_array.squeeze()
|
| 77 |
+
cropped_boxed_tensor = torch.Tensor(cropped_boxed_array)
|
| 78 |
+
prediction, image_transformed = Model_Class.load_and_classify_image(cropped_boxed_tensor)
|
| 79 |
+
|
| 80 |
+
|
| 81 |
+
gradcam = Model_Class.make_GradCAM(image_transformed)
|
| 82 |
+
|
| 83 |
+
nr_axSpA_prob = float(prediction[0].item())
|
| 84 |
+
r_axSpA_prob = float(prediction[1].item())
|
| 85 |
+
|
| 86 |
+
# Decision based on the threshold
|
| 87 |
+
considered = "be considered r-axSpA" if r_axSpA_prob > 0.59 else "not be considered r-axSpA"
|
| 88 |
+
|
| 89 |
+
explanation = f"According to the pre-determined cut-off threshold of 0.59, the image should {considered}. This Tool is for research purposes only."
|
| 90 |
+
|
| 91 |
+
pred_dict = {"nr-axSpA": nr_axSpA_prob, "r-axSpA": r_axSpA_prob}
|
| 92 |
+
|
| 93 |
+
return overlay_image_np, pred_dict, explanation, gradcam, cropped_boxed_array_disp
|
| 94 |
+
|
| 95 |
+
|
| 96 |
+
|
| 97 |
+
|
| 98 |
+
with gr.Blocks(css=css, title="Anatomy Aware axSpA") as iface:
|
| 99 |
+
|
| 100 |
+
gr.Markdown("# Anatomy-Aware Image Classification for radiographic axSpA")
|
| 101 |
+
gr.Markdown(description_markdown, elem_classes="markdown-block")
|
| 102 |
+
|
| 103 |
+
with gr.Row():
|
| 104 |
+
with gr.Column():
|
| 105 |
+
|
| 106 |
+
with gr.Tab("PNG/JPG"):
|
| 107 |
+
input_image = gr.Image(type='filepath', label="Upload an X-ray Image")
|
| 108 |
+
|
| 109 |
+
with gr.Tab("NIfTI/DICOM"):
|
| 110 |
+
input_file = gr.File(type='filepath', label="Upload an X-ray Image")
|
| 111 |
+
|
| 112 |
+
with gr.Row():
|
| 113 |
+
submit_button = gr.Button("Run", variant="primary")
|
| 114 |
+
clear_button = gr.ClearButton()
|
| 115 |
+
|
| 116 |
+
with gr.Column():
|
| 117 |
+
overlay_image_np = gr.Image(label="Segmentation Mask")
|
| 118 |
+
|
| 119 |
+
pred_dict = gr.Label(label="Prediction")
|
| 120 |
+
explanation= gr.Textbox(label="Classification Decision")
|
| 121 |
+
|
| 122 |
+
with gr.Accordion("Additional Information", open=False):
|
| 123 |
+
gradcam = gr.Image(label="GradCAM")
|
| 124 |
+
cropped_boxed_array_disp = gr.Image(label="Bounding Box")
|
| 125 |
+
|
| 126 |
+
submit_button.click(predict_image, inputs = [input_image, input_file], outputs=[overlay_image_np, pred_dict, explanation, gradcam, cropped_boxed_array_disp])
|
| 127 |
+
clear_button.add([input_image,overlay_image_np, pred_dict, explanation, gradcam, cropped_boxed_array_disp])
|
| 128 |
+
gr.HTML(article_html)
|
| 129 |
+
|
| 130 |
+
|
| 131 |
+
if __name__ == "__main__":
|
| 132 |
+
iface.queue()
|
| 133 |
+
iface.launch(server_name='0.0.0.0', server_port=8080)
|
requirements_small.txt
ADDED
|
@@ -0,0 +1,12 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
gradio==4.29.0
|
| 2 |
+
spaces==0.28.3
|
| 3 |
+
numpy==1.22.2
|
| 4 |
+
torch==1.13.0
|
| 5 |
+
torchvision==0.14.0
|
| 6 |
+
scikit-image==0.19.3
|
| 7 |
+
pytorch-lightning==1.8.6
|
| 8 |
+
monai-weekly==1.2.dev2320
|
| 9 |
+
simpleitk==2.2.1
|
| 10 |
+
nibabel==5.2.1
|
| 11 |
+
itk==5.3.0
|
| 12 |
+
grad-cam==1.4.6
|
utils.py
ADDED
|
@@ -0,0 +1,353 @@
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| 1 |
+
from monai.transforms import Transform
|
| 2 |
+
import torch
|
| 3 |
+
import skimage
|
| 4 |
+
import torch
|
| 5 |
+
import SimpleITK as sitk
|
| 6 |
+
import numpy as np
|
| 7 |
+
from PIL import Image
|
| 8 |
+
from io import BytesIO
|
| 9 |
+
import matplotlib.pyplot as plt
|
| 10 |
+
import SimpleITK as sitk
|
| 11 |
+
from matplotlib.colors import ListedColormap
|
| 12 |
+
import base64
|
| 13 |
+
import numpy as np
|
| 14 |
+
from cv2 import dilate
|
| 15 |
+
from scipy.ndimage import label
|
| 16 |
+
|
| 17 |
+
def image_to_base64(image_path):
|
| 18 |
+
with open(image_path, "rb") as image_file:
|
| 19 |
+
return base64.b64encode(image_file.read()).decode('utf-8')
|
| 20 |
+
|
| 21 |
+
class CustomCLAHE(Transform):
|
| 22 |
+
"""Implements Contrast-Limited Adaptive Histogram Equalization (CLAHE) as a custom transform, as described by Qiu et al.
|
| 23 |
+
|
| 24 |
+
Attributes:
|
| 25 |
+
p1 (float): Weighting factor, determines degree of of contour enhacement. Default is 0.6.
|
| 26 |
+
p2 (None or int): Kernel size for adaptive histogram. Default is None.
|
| 27 |
+
p3 (float): Clip limit for histogram equalization. Default is 0.01.
|
| 28 |
+
|
| 29 |
+
"""
|
| 30 |
+
|
| 31 |
+
def __init__(self, p1=0.6, p2=None, p3=0.01):
|
| 32 |
+
self.p1 = p1
|
| 33 |
+
self.p2 = p2
|
| 34 |
+
self.p3 = p3
|
| 35 |
+
|
| 36 |
+
def __call__(self, data):
|
| 37 |
+
"""Apply the CLAHE algorithm to input data.
|
| 38 |
+
|
| 39 |
+
Args:
|
| 40 |
+
data (Union[dict, np.ndarray]): Input data. Could be a dictionary containing the image or the image array itself.
|
| 41 |
+
|
| 42 |
+
Returns:
|
| 43 |
+
torch.Tensor: Transformed data.
|
| 44 |
+
"""
|
| 45 |
+
|
| 46 |
+
if isinstance(data, dict):
|
| 47 |
+
im = data["image"]
|
| 48 |
+
|
| 49 |
+
else:
|
| 50 |
+
im = data
|
| 51 |
+
im = im.numpy()
|
| 52 |
+
|
| 53 |
+
|
| 54 |
+
# remove the first dimension
|
| 55 |
+
im = im[0]
|
| 56 |
+
im = im[None, :, :]
|
| 57 |
+
#im = np.expand_dims(im, axis=0)
|
| 58 |
+
im = skimage.exposure.rescale_intensity(im, in_range="image", out_range=(0, 1))
|
| 59 |
+
im_noi = skimage.filters.median(im)
|
| 60 |
+
im_fil = im_noi - self.p1 * skimage.filters.gaussian(im_noi, sigma=1)
|
| 61 |
+
im_fil = skimage.exposure.rescale_intensity(im_fil, in_range="image", out_range=(0, 1))
|
| 62 |
+
im_ce = skimage.exposure.equalize_adapthist(im_fil, kernel_size=self.p2, clip_limit=self.p3)
|
| 63 |
+
if isinstance(data, dict):
|
| 64 |
+
data["image"] = torch.Tensor(im_ce)
|
| 65 |
+
else:
|
| 66 |
+
data = torch.Tensor(im_ce)
|
| 67 |
+
|
| 68 |
+
return data
|
| 69 |
+
|
| 70 |
+
|
| 71 |
+
|
| 72 |
+
def custom_colormap():
|
| 73 |
+
|
| 74 |
+
cdict = [(0, 0, 0, 0), # Class 0 - fully transparent (background)
|
| 75 |
+
(0, 1, 0, 0.5), # Class 1 - Green with 50% transparency
|
| 76 |
+
(1, 0, 0, 0.5), # Class 2 - Red with 50% transparency
|
| 77 |
+
(1, 1, 0, 0.5)] # Class 3 - Yellow with 50% transparency
|
| 78 |
+
cmap = ListedColormap(cdict)
|
| 79 |
+
return cmap
|
| 80 |
+
|
| 81 |
+
def read_image(image_path):
|
| 82 |
+
try:
|
| 83 |
+
original_image = Image.open(image_path).convert('L')
|
| 84 |
+
original_image_np = np.array(original_image)
|
| 85 |
+
return original_image_np.squeeze()
|
| 86 |
+
|
| 87 |
+
except Exception as e:
|
| 88 |
+
try :
|
| 89 |
+
original_image = sitk.ReadImage(image_path)
|
| 90 |
+
original_image_np = sitk.GetArrayFromImage(original_image)
|
| 91 |
+
return original_image_np.squeeze()
|
| 92 |
+
except Exception as e:
|
| 93 |
+
print("Failed Loading the Image: ", e)
|
| 94 |
+
return None
|
| 95 |
+
|
| 96 |
+
def overlay_mask(image_path, image_mask):
|
| 97 |
+
original_image_np = read_image(image_path).squeeze().astype(np.uint8)
|
| 98 |
+
|
| 99 |
+
#adjust mask intensities for display
|
| 100 |
+
image_mask_disp = image_mask
|
| 101 |
+
plt.figure(figsize=(10, 10))
|
| 102 |
+
plt.imshow(original_image_np, cmap='gray')
|
| 103 |
+
|
| 104 |
+
plt.imshow(image_mask_disp, cmap=custom_colormap(), alpha=0.5)
|
| 105 |
+
plt.axis('off')
|
| 106 |
+
|
| 107 |
+
# Save the overlay to a buffer
|
| 108 |
+
buffer = BytesIO()
|
| 109 |
+
plt.savefig(buffer, format='png', bbox_inches='tight', pad_inches=0)
|
| 110 |
+
buffer.seek(0)
|
| 111 |
+
overlay_image_np = np.array(Image.open(buffer))
|
| 112 |
+
return overlay_image_np, original_image_np
|
| 113 |
+
|
| 114 |
+
|
| 115 |
+
def bounding_box_mask(image, label):
|
| 116 |
+
"""Generates a bounding box mask around a labeled region in an image
|
| 117 |
+
|
| 118 |
+
Args:
|
| 119 |
+
image (SimpleITK.Image): The input image.
|
| 120 |
+
label (SimpleITK.Image): The labeled image containing the region of interest.
|
| 121 |
+
|
| 122 |
+
Returns:
|
| 123 |
+
SimpleITK.Image: An image containing the with the bounding box mask applied with the
|
| 124 |
+
same spacing as the original image.
|
| 125 |
+
|
| 126 |
+
Note:
|
| 127 |
+
This function assumes that the input image and label are SimpleITK.Image objects.
|
| 128 |
+
The returned bounding box mask is a binary image where pixels inside the bounding box
|
| 129 |
+
are set to 1 and others are set to 0.
|
| 130 |
+
"""
|
| 131 |
+
# get original spacing
|
| 132 |
+
original_spacing = image.GetSpacing()
|
| 133 |
+
|
| 134 |
+
# convert image and label to arrays
|
| 135 |
+
image_array = sitk.GetArrayFromImage(image)
|
| 136 |
+
image_array = np.squeeze(image_array)
|
| 137 |
+
label_array = sitk.GetArrayFromImage(label)
|
| 138 |
+
label_array = np.squeeze(label_array)
|
| 139 |
+
|
| 140 |
+
# determine corners of the bounding box
|
| 141 |
+
first_nonzero_row_index = np.nonzero(np.any(label_array != 0, axis=1))[0][0]
|
| 142 |
+
last_nonzero_row_index = np.max(np.nonzero(np.any(label_array != 0, axis=1)))
|
| 143 |
+
first_nonzero_column_index = np.nonzero(np.any(label_array != 0, axis=0))[0][0]
|
| 144 |
+
last_nonzero_column_index = np.max(np.nonzero(np.any(label_array != 0, axis=0)))
|
| 145 |
+
|
| 146 |
+
top_left_corner = (first_nonzero_row_index, first_nonzero_column_index)
|
| 147 |
+
bottom_right_corner = (last_nonzero_row_index, last_nonzero_column_index)
|
| 148 |
+
|
| 149 |
+
# define the bounding box as an array mask
|
| 150 |
+
bounding_box_array = label_array.copy()
|
| 151 |
+
bounding_box_array[
|
| 152 |
+
top_left_corner[0] : bottom_right_corner[0] + 1,
|
| 153 |
+
top_left_corner[1] : bottom_right_corner[1] + 1,
|
| 154 |
+
] = 1
|
| 155 |
+
|
| 156 |
+
# add channel dimension
|
| 157 |
+
bounding_box_array = bounding_box_array[None, ...].astype(np.uint8)
|
| 158 |
+
|
| 159 |
+
# get Image from Array Mask and apply original spacing
|
| 160 |
+
bounding_box_image = sitk.GetImageFromArray(bounding_box_array)
|
| 161 |
+
bounding_box_image.SetSpacing(original_spacing)
|
| 162 |
+
return bounding_box_image
|
| 163 |
+
|
| 164 |
+
|
| 165 |
+
def threshold_based_crop(image):
|
| 166 |
+
"""
|
| 167 |
+
Use Otsu's threshold estimator to separate background and foreground. In medical imaging the background is
|
| 168 |
+
usually air. Then crop the image using the foreground's axis aligned bounding box.
|
| 169 |
+
Args:
|
| 170 |
+
image (SimpleITK image): An image where the anatomy and background intensities form a
|
| 171 |
+
bi-modal distribution
|
| 172 |
+
(the assumption underlying Otsu's method.)
|
| 173 |
+
Return:
|
| 174 |
+
Cropped image based on foreground's axis aligned bounding box.
|
| 175 |
+
"""
|
| 176 |
+
|
| 177 |
+
inside_value = 0
|
| 178 |
+
outside_value = 255
|
| 179 |
+
label_shape_filter = sitk.LabelShapeStatisticsImageFilter()
|
| 180 |
+
# uncomment for debugging
|
| 181 |
+
#sitk.WriteImage(image, "./image.png")
|
| 182 |
+
label_shape_filter.Execute(sitk.OtsuThreshold(image, inside_value, outside_value))
|
| 183 |
+
bounding_box = label_shape_filter.GetBoundingBox(outside_value)
|
| 184 |
+
return sitk.RegionOfInterest(
|
| 185 |
+
image,
|
| 186 |
+
bounding_box[int(len(bounding_box) / 2) :],
|
| 187 |
+
bounding_box[0 : int(len(bounding_box) / 2)],
|
| 188 |
+
)
|
| 189 |
+
|
| 190 |
+
def creat_SIJ_mask(image, input_label):
|
| 191 |
+
"""
|
| 192 |
+
Create a mask for the sacroiliac joints (SIJ) from pelvis and sascrum segmentation mask
|
| 193 |
+
|
| 194 |
+
Args:
|
| 195 |
+
image (SimpleITK.Image): x-ray image.
|
| 196 |
+
input_label (SimpleITK.Image): Segmentation mask containing labels for sacrum, left- and right pelvis
|
| 197 |
+
|
| 198 |
+
Returns:
|
| 199 |
+
SimpleITK.Image: Mask of the SIJ
|
| 200 |
+
|
| 201 |
+
"""
|
| 202 |
+
|
| 203 |
+
original_spacing = image.GetSpacing()
|
| 204 |
+
# uncomment for debugging
|
| 205 |
+
#sitk.WriteImage(input_label, "./input_label.png")
|
| 206 |
+
mask_array = sitk.GetArrayFromImage(input_label).squeeze()
|
| 207 |
+
|
| 208 |
+
sacrum_value = 1
|
| 209 |
+
left_pelvis_value = 2
|
| 210 |
+
right_pelvis_value = 3
|
| 211 |
+
background_value = 0
|
| 212 |
+
|
| 213 |
+
|
| 214 |
+
sacrum_mask = (mask_array == sacrum_value)
|
| 215 |
+
|
| 216 |
+
first_nonzero_column_index = np.nonzero(np.any(sacrum_mask != 0, axis=0))[0][0]
|
| 217 |
+
last_nonzero_column_index = np.max(np.nonzero(np.any(sacrum_mask != 0, axis=0)))
|
| 218 |
+
box_width=last_nonzero_column_index-first_nonzero_column_index
|
| 219 |
+
|
| 220 |
+
dilation_extent = int(np.round(0.05 * box_width))
|
| 221 |
+
|
| 222 |
+
dilated_sacrum_mask = dilate_mask(sacrum_mask, dilation_extent)
|
| 223 |
+
|
| 224 |
+
intersection_left = (dilated_sacrum_mask & (mask_array == left_pelvis_value))
|
| 225 |
+
if np.all(intersection_left == 0):
|
| 226 |
+
print("Warning: No left intersection")
|
| 227 |
+
left_pelvis_mask = (mask_array == 2)
|
| 228 |
+
intersection_left = create_median_height_array(left_pelvis_mask)
|
| 229 |
+
|
| 230 |
+
intersection_left = keep_largest_component(intersection_left)
|
| 231 |
+
|
| 232 |
+
intersection_right = (dilated_sacrum_mask & (mask_array == right_pelvis_value))
|
| 233 |
+
if np.all(intersection_right == 0):
|
| 234 |
+
print("Warning: No right intersection")
|
| 235 |
+
right_pelvis_mask = (mask_array == 3)
|
| 236 |
+
intersection_right = create_median_height_array(right_pelvis_mask)
|
| 237 |
+
intersection_right = keep_largest_component(intersection_right)
|
| 238 |
+
|
| 239 |
+
intersection_mask = intersection_left +intersection_right
|
| 240 |
+
intersection_mask = intersection_mask[None, ...]
|
| 241 |
+
|
| 242 |
+
instersection_mask_im = sitk.GetImageFromArray(intersection_mask)
|
| 243 |
+
instersection_mask_im.SetSpacing(original_spacing)
|
| 244 |
+
return instersection_mask_im
|
| 245 |
+
|
| 246 |
+
def dilate_mask(mask, extent):
|
| 247 |
+
"""
|
| 248 |
+
Keeps only the largest connected component in a binary segmentation mask.
|
| 249 |
+
|
| 250 |
+
Args:
|
| 251 |
+
mask (numpy.ndarray): A numpy array representing the binary segmentation mask,
|
| 252 |
+
with 1s indicating the label and 0s indicating the background.
|
| 253 |
+
|
| 254 |
+
Returns:
|
| 255 |
+
numpy.ndarray: A modified version of the input mask, where only the largest
|
| 256 |
+
connected component is retained, and other components are set to 0.
|
| 257 |
+
|
| 258 |
+
"""
|
| 259 |
+
mask_uint8 = mask.astype(np.uint8)
|
| 260 |
+
|
| 261 |
+
kernel = np.ones((2*extent+1, 2*extent+1), np.uint8)
|
| 262 |
+
dilated_mask = dilate(mask_uint8, kernel, iterations=1)
|
| 263 |
+
return dilated_mask
|
| 264 |
+
|
| 265 |
+
def mask_and_crop(image, input_label):
|
| 266 |
+
"""
|
| 267 |
+
Performs masking and cropping operations on an image and its label.
|
| 268 |
+
|
| 269 |
+
Args:
|
| 270 |
+
image (SimpleITK.Image): The image to be processed.
|
| 271 |
+
label (SimpleITK.Image): The corresponding label image.
|
| 272 |
+
|
| 273 |
+
Returns:
|
| 274 |
+
tuple: A tuple containing two SimpleITK.Image objects.
|
| 275 |
+
- cropped_boxed_image: The image after applying bounding box masking and cropping.
|
| 276 |
+
- mask: The binary mask corresponding to the label after cropping.
|
| 277 |
+
|
| 278 |
+
Note:
|
| 279 |
+
This function relies on other functions: bounding_box_mask() and threshold_based_crop().
|
| 280 |
+
"""
|
| 281 |
+
input_label = creat_SIJ_mask(image,input_label)
|
| 282 |
+
box_mask = bounding_box_mask(image, input_label)
|
| 283 |
+
|
| 284 |
+
boxed_image = sitk.Mask(image, box_mask, maskingValue=0, outsideValue=0)
|
| 285 |
+
masked_image = sitk.Mask(image, input_label, maskingValue=0, outsideValue=0)
|
| 286 |
+
|
| 287 |
+
cropped_boxed_image = threshold_based_crop(boxed_image)
|
| 288 |
+
cropped_masked_image = threshold_based_crop(masked_image)
|
| 289 |
+
|
| 290 |
+
mask = np.squeeze(sitk.GetArrayFromImage(cropped_masked_image))
|
| 291 |
+
mask = np.where(mask > 0, 1, 0)
|
| 292 |
+
mask = sitk.GetImageFromArray(mask[None, ...])
|
| 293 |
+
return cropped_boxed_image, mask
|
| 294 |
+
|
| 295 |
+
def create_median_height_array(mask):
|
| 296 |
+
"""
|
| 297 |
+
Creates an array based on the median height of non-zero elements in each column of the input mask.
|
| 298 |
+
|
| 299 |
+
Args:
|
| 300 |
+
mask (numpy.ndarray): A binary mask with 1s representing the label and 0s the background.
|
| 301 |
+
|
| 302 |
+
Returns:
|
| 303 |
+
numpy.ndarray: A new binary mask array with columns filled based on the median height,
|
| 304 |
+
or None if the input mask has no non-zero columns.
|
| 305 |
+
|
| 306 |
+
Note:
|
| 307 |
+
This function is only used when there is no intersection between pelvis and sacrum, and creates an alternative
|
| 308 |
+
SIJ mask, that serves as an approximate replacement.
|
| 309 |
+
"""
|
| 310 |
+
rows, cols = mask.shape
|
| 311 |
+
column_details = []
|
| 312 |
+
|
| 313 |
+
for col in range(cols):
|
| 314 |
+
column_data = mask[:, col]
|
| 315 |
+
non_zero_indices = np.nonzero(column_data)[0]
|
| 316 |
+
if non_zero_indices.size > 0:
|
| 317 |
+
height = non_zero_indices[-1] - non_zero_indices[0] + 1
|
| 318 |
+
start_idx = non_zero_indices[0]
|
| 319 |
+
column_details.append((height, start_idx, col))
|
| 320 |
+
|
| 321 |
+
if not column_details:
|
| 322 |
+
return None
|
| 323 |
+
median_height = round(np.median([h[0] for h in column_details]))
|
| 324 |
+
median_cols = [(col, start_idx) for height, start_idx, col in column_details if height == median_height]
|
| 325 |
+
new_array = np.zeros_like(mask, dtype=int)
|
| 326 |
+
for col, start_idx in median_cols:
|
| 327 |
+
start_col = max(0, col - 5)
|
| 328 |
+
end_col = min(cols, col + 5)
|
| 329 |
+
new_array[start_idx:start_idx + median_height, start_col:end_col] = 1
|
| 330 |
+
return new_array
|
| 331 |
+
|
| 332 |
+
def keep_largest_component(mask):
|
| 333 |
+
"""
|
| 334 |
+
Identifies and retains the largest connected component in a binary segmentation mask.
|
| 335 |
+
|
| 336 |
+
Args:
|
| 337 |
+
mask (numpy.ndarray): A binary mask with 1s representing the label and 0s the background.
|
| 338 |
+
|
| 339 |
+
Returns:
|
| 340 |
+
numpy.ndarray: The modified mask with only the largest connected component.
|
| 341 |
+
"""
|
| 342 |
+
# Label the connected components
|
| 343 |
+
labeled_array, num_features = label(mask)
|
| 344 |
+
|
| 345 |
+
# If no features are found, return the original mask
|
| 346 |
+
if num_features <= 1:
|
| 347 |
+
return mask
|
| 348 |
+
|
| 349 |
+
# Find the largest connected component
|
| 350 |
+
largest_component = np.argmax(np.bincount(labeled_array.flat)[1:]) + 1
|
| 351 |
+
|
| 352 |
+
# Generate the mask for the largest component
|
| 353 |
+
return (labeled_array == largest_component).astype(mask.dtype)
|