Datasets:
license: cc-by-4.0
task_categories:
- visual-question-answering
- zero-shot-image-classification
- zero-shot-object-detection
tags:
- biology
- organism
- fish
- bird
- butterfly
- image classification
- zero-shot
- traits
- trait-detection
- vlms
- benchmarks
- CV
language:
- en
pretty_name: VLM4Bio
size_categories:
- 10K<n<100K
configs:
- config_name: Fish
data_files:
- split: species_classification
path: datasets/Fish/metadata/metadata_10k.csv
- split: species_classification_easy
path: datasets/Fish/metadata/metadata_easy.csv
- split: species_classification_medium
path: datasets/Fish/metadata/metadata_medium.csv
- split: species_classification_prompting
path: datasets/Fish/metadata/metadata_prompting.csv
- config_name: Bird
data_files:
- split: species_classification
path: datasets/Bird/metadata/metadata_10k.csv
- split: species_classification_easy
path: datasets/Bird/metadata/metadata_easy.csv
- split: species_classification_medium
path: datasets/Bird/metadata/metadata_medium.csv
- split: species_classification_prompting
path: datasets/Bird/metadata/metadata_prompting.csv
- config_name: Butterfly
data_files:
- split: species_classification
path: datasets/Butterfly/metadata/metadata_10k.csv
- split: species_classification_easy
path: datasets/Butterfly/metadata/metadata_easy.csv
- split: species_classification_medium
path: datasets/Butterfly/metadata/metadata_medium.csv
- split: species_classification_hard
path: datasets/Butterfly/metadata/metadata_hard.csv
- split: species_classification_prompting
path: datasets/Butterfly/metadata/metadata_prompting.csv
Dataset Card for VLM4Bio
Dataset Details
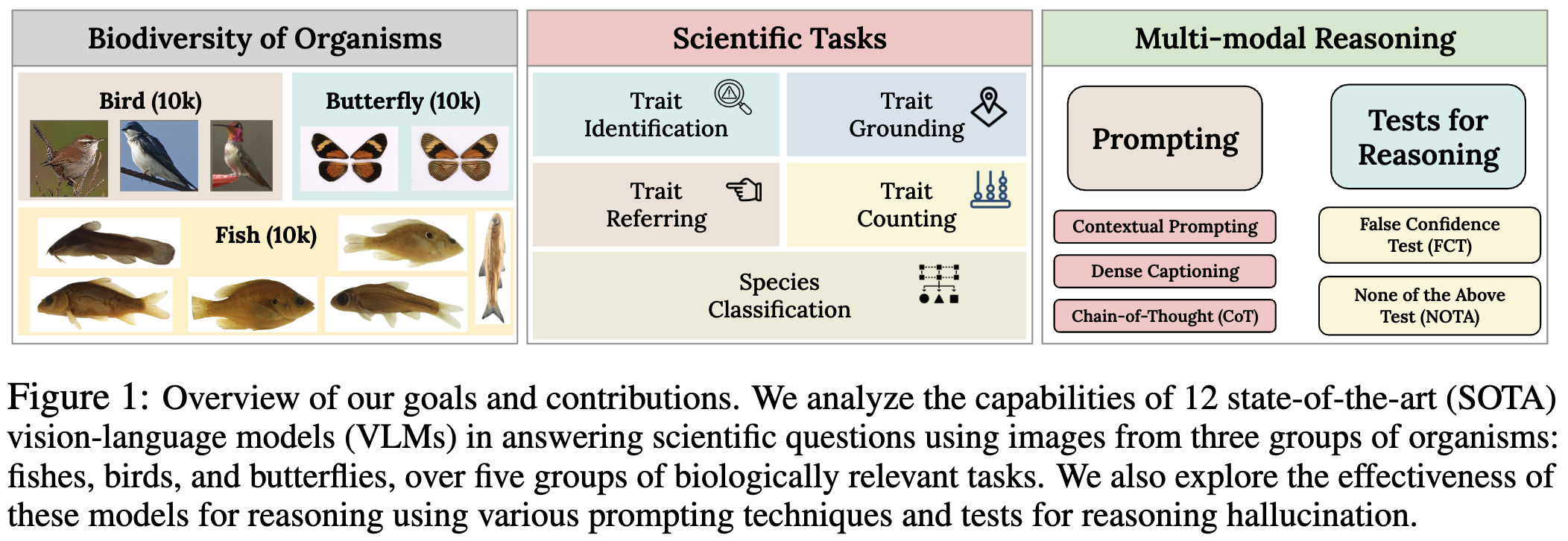
VLM4Bio is a benchmark dataset of scientific question-answer pairs used to evaluate pretrained VLMs for trait discovery from biological images. VLM4Bio consists of images of three taxonomic groups of organisms: fish, birds, and butterflies, each containing around 10k images.
- Repository: VLM4Bio GitHub
- Paper: arXiv
Dataset Description
VLM4Bio is a large, annotated dataset, consisting of 469K question-answer pairs involving around 30K images from three groups of organisms: fish, birds, and butterflies, covering five biologically relevant tasks. The scientifically relevant tasks in organismal biology includes species classification, trait identification, trait grounding, trait referring, and trait counting. These tasks are designed to test different facets of VLM performance in organismal biology, ranging from measuring predictive accuracy to assessing their ability to reason about their predictions using visual cues of known biological traits. For example, the tasks of species classification test the ability of VLMs to discriminate between species, while in trait grounding and referring, we specifically test if VLMs are able to localize morphological traits (e.g., the presence of fins of fish or patterns and colors of birds) within the image. We consider two types of questions in this dataset. First, we consider open-ended questions, where we do not provide any answer choices (or options) to the VLM in the input prompt. The second type is multiple-choice (MC) questions, where we provide four choices of candidate answers for the VLM to choose from (out of which only one is correct while the remaining three are randomly selected from the set of all possible answers).
Supported Tasks and Leaderboards
The following figure illustrates VLM4Bio tasks with different question types.
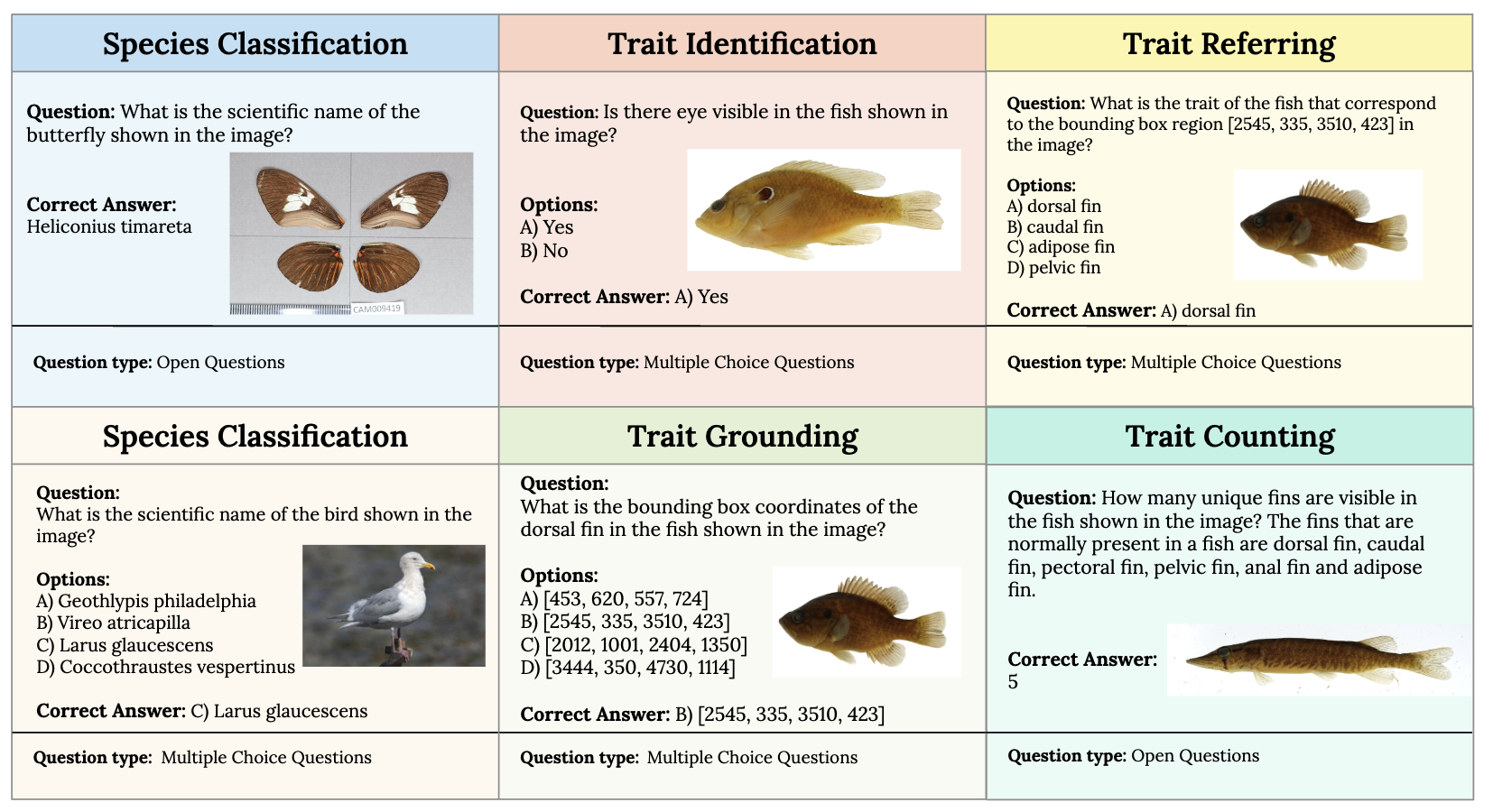
The following table demonstrates the leaderboard of the VLM baselines in terms of zero-shot accuracy.
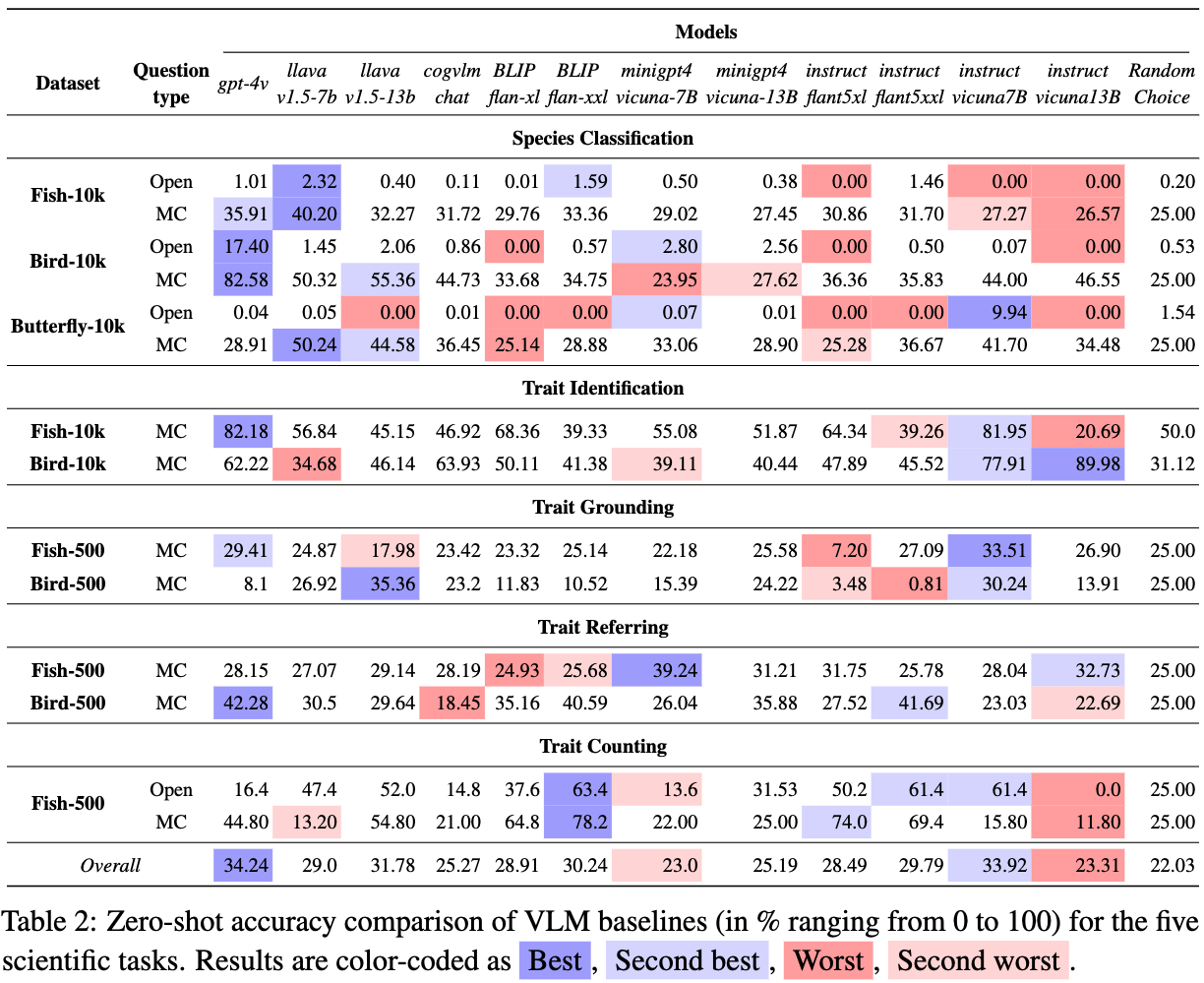
Languages
English, Latin
Dataset Structure
Instructions for downloading the dataset
Data Instances
Data Fields
Fish Files:
identification_imagelist_10k.txt:identification_matrix.csv:imagelist_10k.txt: List of image filenames for species classification and trait identification.imagelist_500.txt: 500 image subset ofimagelist_10k.txtfor trait detection and counting.metadata_10k.csv: Image filenames (fileNameAsDelivered, unique identifier) paired with their respective scientific names (scientificName).metadata_500.csv: 500 image subset ofmetadata_10k.csvfor trait detection and counting.ARKIDis unique identifier from Fish-AIR, links to full metadata information infull_fish_metadata.csv.processed_identification_imagelist_10k.txt: List of images included in theprocessed_identification_matrix.csv.processed_identification_matrix.csv: Presence/Absence indicator for 10 external (visible) traits:eye,head,mouth,barbel,dorsal fin,two dorsal fins,adipose fin,pectoral fin,pelvic fin,anal fin. Unique identifier is thefileNameAsDelivered, and scientific name is indicated (scientificName).
Bird Files:
bird_imagelist_10k.txt: List of image filenames for species classification and trait identification.bird_metadata_10k.csv: Image filenames (fileNameAsDelivered, unique identifier) paired with their respective scientific names (scientificName).identification.csv:processed_identification.csv:trait_category_map.pkl:
Butterfly Files:
imagelist.txt: List of image filenames for species classification.metadata.csv: Image filenames (fileNameAsDelivered, unique identifier) paired with their respective scientific names (scientificName).
VLM Prompts are determined through code available in the GitHub Repository; they are summarized in the task diagram above.
Data Splits
These images were all used for benchmarking current state-of-the-art VLMs on biological tasks.
Curation Rationale
Source Data
We collected images of three taxonomic groups of organisms: fish, birds, and butterflies, each containing around 10k images.
Fish
Images for fish (Fish-10k) were curated from the larger image collection, Fish-AIR, which contains images from the Great Lakes Invasives Network (GLIN) and Integrated Digitized Biocollections (iDigBio).
These images originate from various museum collections, including the following:
- Illinois Natural History Survey (INHS)
- Minnesota Biodiversity Atlas, Bell Museum
- University of Michigan Museum of Zoology (UMMZ), Division of Fishes
- University of Wisconsin-Madison Zoological Museum - Fish
- Field Museum of Natural History (Zoology, FMNH) Fish Collection
- The Ohio State University Fish Division, Museum of Biological Diversity (OSUM), Occurrence dataset
Phenoscape and FishBase were used to obtain the information on traits.
Data Processing:
We created the Fish-10k dataset by randomly sampling 10K images and preprocessing the images to crop and remove the background. For consistency, we leverage GroundingDINO to crop the fish body from the background and Segment Anything Model (SAM) to remove the background. This is the same processing done in Fish-Vista, more details and the code is available here.
Bird
We create the Bird-10k dataset from the CUB-200-2011 dataset. We obtain the scientific names from the iNatLoc dataset.
Data Processing:
For Bird-10k, we take 190 species for which the common name to scientific name mapping is available. This results in a fairly balanced dataset. Please download the images following the directions under Dataset Structure.
Butterflies
We created the Butterfly-10k dataset from the Heliconius Collection (Cambridege Butterfly) dataset.
Data Processing:
For the Butterfly-10k, we carefully sampled 10K images from the Heliconius Collection dataset to ensure the images capture unique specimens and represent a diverse set of species. We adopt the following steps:
- We filter out images with more than one image from the same view (i.e., dorsal or ventral).
- We ensure each species has a minimum of 20 images and no more than 2,000 images.
Annotations
Scientific Names
The scientific names for the images of Fish-10k and Butterfly-10k were obtained directly from their respective sources.
For Bird-10k, we obtained the scientific names from the iNatLoc dataset.
In total, we curated around 31K question-answer pairs in both open and multiple-choice (MC) question-formats for evaluating species classification tasks.
Trait information
The species-level trait presence/absence matrix for Fish-10k was manually curated with the help of biological experts co-authored in this paper. We leveraged the Phenoscape knowledge base and FishBase along with manual annotations to procure the presence-absence trait information. We constructed approximately xK question-answer pairs for Fish-10k
For Bird-10k, we obtained the trait matrix from the attribute annotations provided along with CUB-200-2011.
In total, we constructed approximately 380K question-answer pairs for trait identification tasks.
Grounding and referring
For grounding and referring VQA tasks, the ground truths were manually annotated with the help of expert biologists on our team. We manually annotated bounding boxes corresponding to the traits of 500 fish specimens and 500 bird specimens, which are subsets of the larger Fish-10k and Bird-10k datasets, respectively. We used the CVAT tool for annotation.
Personal and Sensitive Information
None
Considerations for Using the Data
The fish-10 K and Butterfly-10K datasets are not balanced for the species classification task, while the bird-10 K dataset is balanced. Since the fish images are collected from different museums, they may inherit a small bias.
Licensing Information
This dataset (the compilation) has been licensed under CC BY 4.0. However, images may be licensed under different terms (as noted above). For license and citation information by image, see our license file.
- The fish images are from iDigBio and GLIN, whose metadata and source URLs were accessed through Fish-AIR. All iDigBio images are in the public domain (CC0). The GLIN images are all CC BY-NC.
- All the bird images are sourced from the CUB-200-2011 dataset; CalTech indicates that they do not own the copyrights to these images, and that their use is restricted to non-commercial research and educational purposes.
- All butterfly images are from the Butterfly Genetics Group at University of Cambridge and are licensed under Creative Commons Attribution 4.0 International.
Each image in this dataset is provided under the least restrictive terms allowed by its licensing requirements as provided to us (i.e., we impose no additional restrictions past those specified by licenses in the license file).
We provide licensing information for every individual image within the fish and butterfly images in license-metadata/fish-licenses.csv and license-metadata/butterfly-licenses.csv respectively. The source_link and citation for each of the butterfly images can be obtained by matching the record_number field to the record numbers in license-metadata/butterfly-licenses.json.
Citation
Please cite our work as follows:
@misc{maruf2024vlm4bio,
title={VLM4Bio: A Benchmark Dataset to Evaluate Pretrained Vision-Language Models for Trait Discovery from Biological Images},
author={M. Maruf and Arka Daw and Kazi Sajeed Mehrab and Harish Babu Manogaran and Abhilash Neog and Medha Sawhney and Mridul Khurana and James P. Balhoff and Yasin Bakis and Bahadir Altintas and Matthew J. Thompson and Elizabeth G. Campolongo and Josef C. Uyeda and Hilmar Lapp and Henry L. Bart and Paula M. Mabee and Yu Su and Wei-Lun Chao and Charles Stewart and Tanya Berger-Wolf and Wasila Dahdul and Anuj Karpatne},
year={2024},
eprint={2408.16176},
archivePrefix={arXiv},
primaryClass={cs.CV},
url={https://arxiv.org/abs/2408.16176},
}
Please be sure to also cite the original data sources using all of the citations provided in the following:
- Sources for Fish-10k: license-metadata/fish-data-bib.bib.
- Sources for Bird-10k: license-metadata/bird-data-bib.bib
- Sources for Butterfly-10k: license-metadata/butterfly-data-bib.bib
Acknowledgements
This work was supported by the Imageomics Institute, which is funded by the US National Science Foundation's Harnessing the Data Revolution (HDR) program under Award #2118240 (Imageomics: A New Frontier of Biological Information Powered by Knowledge-Guided Machine Learning). Any opinions, findings and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Dataset Card Authors
M. Maruf, Kazi Sajeed Mehrab and Elizabeth G. Campolongo