tags:
- molecular language model
- SELFIES
- molecule optimization
inference: false
MolGen-large-opt
MolGen-large-opt was introduced in the paper "Domain-Agnostic Molecular Generation with Self-feedback" and first released in this repository.
Model description
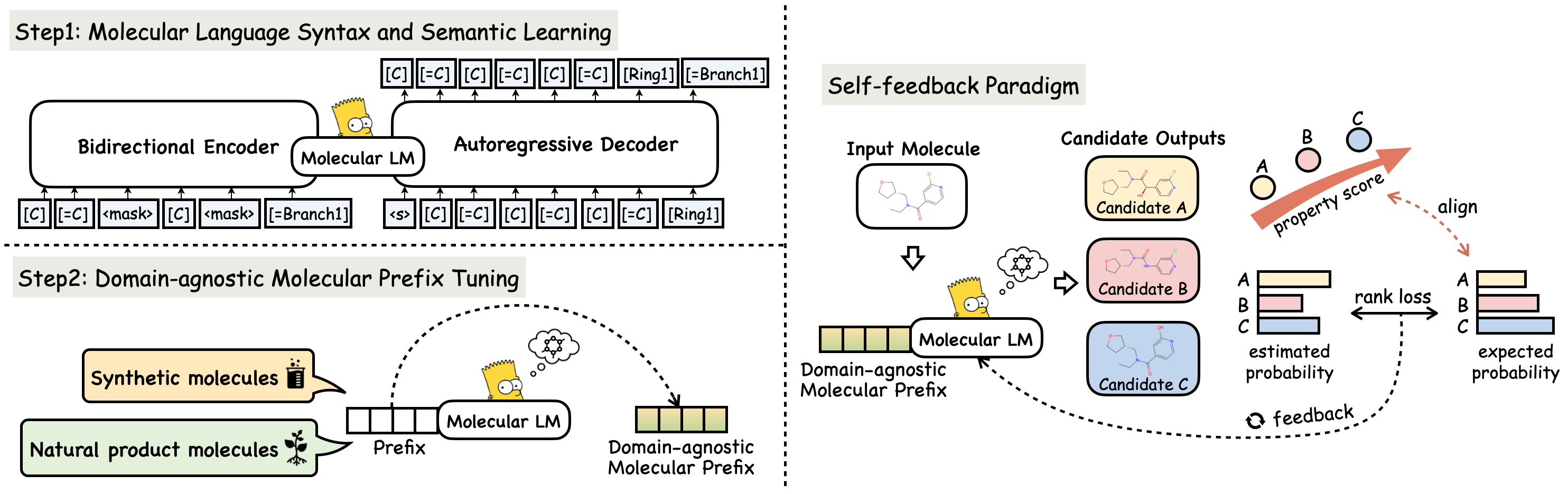
MolGen-large-opt is the fine-tuned version of MolGen-large. MolGen-large is the first pre-trained model that only produces chemically valid molecules. With a training corpus of over 100 million molecules in SELFIES representation, MolGen-large learns the intrinsic structural patterns of molecules by mapping corrupted SELFIES to their original forms. Specifically, MolGen-large employs a bidirectional Transformer as its encoder and an autoregressive Transformer as its decoder. Through its carefully designed multi-task molecular prefix tuning (MPT), MolGen-large-opt can generate molecules with desired properties, making it a valuable tool for molecular optimization.
Intended uses
You can use the fine-tuned model for molecule optimization for downstream tasks. See the repository to look for fine-tune details on a task that interests you.
How to use
Molecule optimization example:
>>> from transformers import AutoTokenizer, AutoModelForSeq2SeqLM
>>> tokenizer = AutoTokenizer.from_pretrained("zjunlp/MolGen-large-opt")
>>> model = AutoModelForSeq2SeqLM.from_pretrained("zjunlp/MolGen-large-opt")
>>> sf_input = tokenizer("[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]", return_tensors="pt")
>>> # beam search
>>> molecules = model.generate(input_ids=sf_input["input_ids"],
attention_mask=sf_input["attention_mask"],
max_length=35,
min_length=5,
num_return_sequences=5,
num_beams=5)
>>> sf_output = [tokenizer.decode(g, skip_special_tokens=True, clean_up_tokenization_spaces=True).replace(" ","") for g in molecules]
['[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]',
'[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]',
'[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]',
'[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]',
'[N][#C][C][C][C@@H1][C][C][C][C][C][C][C][C][C][C][C][C][C][C][Ring1][N][=O]']
BibTeX entry and citation info
@article{fang2023molecular,
title={Molecular Language Model as Multi-task Generator},
author={Fang, Yin and Zhang, Ningyu and Chen, Zhuo and Fan, Xiaohui and Chen, Huajun},
journal={arXiv preprint arXiv:2301.11259},
year={2023}
}